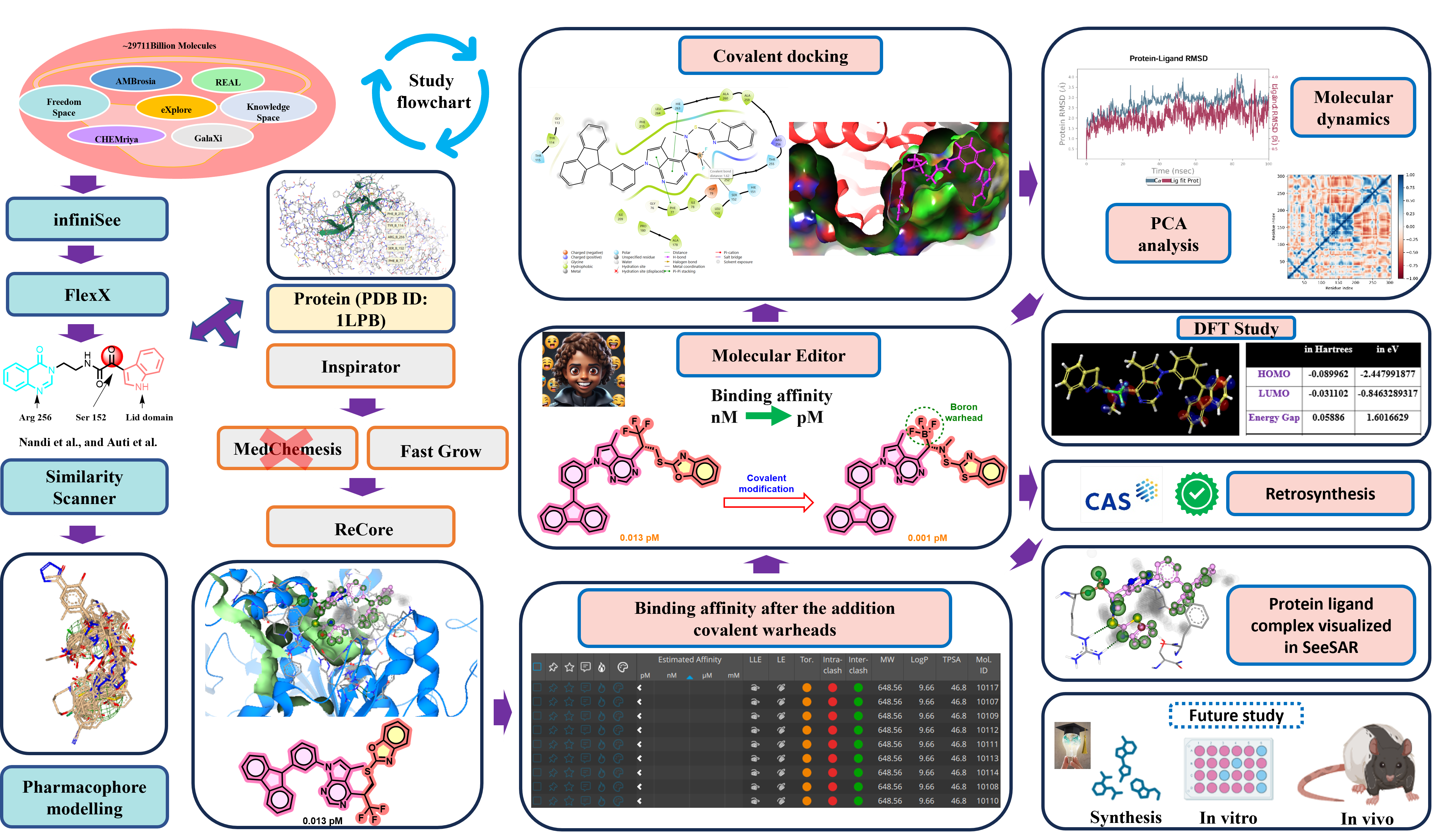
The inhibition of pancreatic lipase (PL) is considered one of the well-accepted strategies in the treatment of obesity (FDA-approved drug Orlistat; β-lactone covalent warhead). In search of a covalent PL inhibitor, during these twelve months of this project, we had successfully found the covalent molecule with a binding affinity of 0.000001 nM from 0.223 nM (quinazolinone-indole hybrid; Nandi et al., 2024; Auti et al., 2022) against the PL enzyme. In infiniSee, the Scaffold Hopper mode was used to obtain a diverse range of compounds. At first, the scaffold hopping of the quinazolinone-indole hybrid lead was performed in the module through the extensive exploration of seven different chemical spaces (AMBrosia, REAL Space, eXplore, Freedom Space, CHEMriya, GalaXi, and literature-based virtual KnowledgeSpace). Optibrium filters were used to attain the better pharmacokinetics. Three visualizations, i.e., scaffold by skeleton, Molecule by scaffold, and skeleton, were used for analysis. These molecules were considered for FlexX for further analysis. After all the screening processes (infiniSee, MedChemesis, and ReCore), ~300 candidates were found to have a 0.013-0.492 pM binding affinity, which is very promising. Optibrium filters were used to attain the better pharmacokinetics. Moreover, parameters like LE, LLE, Torsion quality, and inter- and intra molecular clash were considered for the analysis. Furthermore, the resulting candidates comprised different types of linkers, i.e., e-W, e-D, hydrophobic, and donor-acceptor. However, the linkers with diverse nature, length, and rigidity are desired, for which the further employment of the Molecular Editor mode was used for covalent molecules’ designing. For the purpose of designing covalent molecules, we performed ~70 covalent modifications, which resulted in even better binding affinity (0.001-0.013 pM). Next, covalent docking was performed using the CovDock module, which reported acceptable docking scores (-6 to -8 kcal/mol). Further, molecular dynamics (MD) simulations, induced-fit docking (IFD), and the MM-GBSA study of the protein-ligand complex of the best docking score-containing compound were performed, and the results indicated the complex stability. Moreover, a principal component analysis (PCA) study was performed for examining the dynamics of the conformational changes and the folding of protein when bound to the lead molecule. Additionally, a density-functional theory (DFT) study was performed to see the electronic properties of the lead molecule. Based on the retrosynthetic approach (via CAS SciFinder database), the Suzuki coupling reaction could be used for the synthesis of the lead molecule. Synthetically feasible compounds should be validated by in vitro as well as in vivo studies in future.
After 1 year, Arijit has achieved the following goals:
- In infiniSee, the Scaffold Hopper mode was used to obtain a diverse range of compounds. At first, the scaffold hopping of the quinazolinone-indole hybrid lead was performed in the module through the extensive exploration of seven different chemical spaces. Scaffold by skeleton, Molecule by scaffold and skeleton were used for analysis additionally, these molecules were considered for FlexX for further analysis. Furthermore, molecules with top binding affinity (pM-nM) were taken into similarity scanner and were selected for pharmacophore modelling. In the search for a promising candidate, the Inspirator mode was taken into consideration. First the MedChemesis-Analoging was used; unfortunately, no pose generated for that molecule. Secondly, ReCore-Rescaffolding was used for diverse range of compounds with different scaffold. The binding affinity were drastically changed from nM range to pM range (or 0.013-0.492 pM).
- ReCore-obtained best molecule (0.013 pM) was further optimized in Molecular Editor mode for the covalent warhead by considering Bemis-Murcko scaffold. In Schrodinger, QSAR studies were done for those molecules with their binding affinity values (obtained from SeeSAR). In a search of covalent inhibitors of Ser152 (pancreatic lipase), different modifications of the molecule were done based on “most frequently used covalent warheads” in SeeSAR. Interestingly, after the addition of the covalent warheads such as boron, α-ketoamide, sulfonyl fluoride, aziridine, nitroalkane, urea, and alkynylamine, etc., the binding affinity was decreased to 0.001-0.013 pM. Therefore, all the covalent molecules were considered for covalent docking in Schrodinger (CovDock module). Moreover, after successful covalent docking, all the molecules had a -6 to -8 kcal/mol docking score, which are the covalent docking scores in acceptable ranges.
- After careful consideration of the above-mentioned warheads, it was shown that the boron-containing compounds were the compound with the best covalent docking score (-8.009 Kcal/mol) in Schrodinger as well as in the SeeSAR binding affinity analysis. Furthermore, after analyzing the protein-ligand interaction of the best compound, it was reported that all the required amino acids (i.e., Phe77, Ser152, Phe215, Arg256, and His263) interacted (based on literature available) with the pancreatic lipase enzyme. The IFDScore (-972.03) and MMGBSA (-31.06 Kcal/mol) values are for this molecule. Also, the stability of the protein-ligand complex was optimized by molecular dynamics simulations. The best molecule had ideal pharmacokinetics and druglike properties (optibrium filter-enabled). Furthermore, in finding synthetically feasible methods, the CAS SciFinder database was used, and a low-cost synthetic scheme was obtained with a good atom economy.